Starch synthase (EC 2.4.1.21) is an enzyme that catalyzes the transfer of glucose molecules from ADP-glucose to the non-reducing end of a pre-existing α-(1→4) linked glucan chain, to which the monosaccharides are attached via an α-(1→4) glycosidic bond.[1]
(1→4)-α-D-glucosyl(n) + ADP-α-D-glucose ⇌ (1→4)-α-D-glucosyl(n+1) + ADP + H+
This enzymatic activity is involved in the synthesis of amylose and amylopectin, the two major constituents of starch, which is the main storage form of carbohydrates in plants.[2]
Starch synthase belongs to the family of glycosyltransferases (EC 2.4), like starch phosphorylase (EC 2.4.1.1), another enzyme involved in starch metabolism, and glycogen synthase (EC 2.4.1.11) and glycogen phosphorylase (EC 2.4.1.1), which are involved in glycogen synthesis and glycogen degradation (glycogenolysis), respectively.[3]
However, whereas glycogen synthase uses UDP-glucose as the glucosyl donor, and starch phosphorylase uses glucose 1-phosphate, starch synthase uses ADP-glucose.[4][5]
Contents
- Isoforms
- Starch synthase and MOS
- Starch synthase and amylopectin synthesis
- GBSS and amylose synthesis
- References
Isoforms
Six isoforms of starch synthase are known in plants. They are structurally related proteins, with five involved in the synthesis of amylopectin, referred to as starch synthase I, II, III, IV and V (SSI, SSII, SSIII, SSIV and SSV, respectively), and one involved in the synthesis of amylose, the granule-bound starch synthase (GBSS; EC 2.4.1.242).[3]
GBSS, SSI, SSII, SSIII, and SSIV possess catalytic activity, whereas SSV lacks it.[6]
GBSS is almost exclusively bound to starch granules and is located mainly within them, as evidenced by protease treatment of the granule surface. The other isoforms of starch synthase are found either only in the stroma of plastids or partitioned between starch granules and the stroma, and are referred to as soluble starch synthases.[7]
Starch synthases I, II, III, and IV catalyze the addition of a single glucose residue per substrate encounter, a distributive mode of action, whereas GBSS catalyzes the addition of more than one glucosidic unit per substrate encounter, a processive mode of action.[8]
| Isoform | Polysaccharide synthesized | Mode of action | Location | Notes |
|---|---|---|---|---|
| GBSS | Amylose | Processive: adds multiple glucose units per encounter | Primarily bound to starch granules | Catalyzes the elongation of amylose chains |
| SSI–IV | Amylopectin | Distributive: adds one glucose unit per encounter | Stroma and/or soluble in starch granules | Exhibit catalytic activity; contribute to amylopectin branching |
| SSV | Amylopectin | Not specified | Not specified | Lacks catalytic activity; function not fully understood |
Starch synthase and MOS
Starch synthases involved in the early steps of amylose and amylopectin synthesis require short malto-oligosaccharides (MOS) to initiate the de novo synthesis of these two polysaccharides.
These small oligosaccharides, α-(1→4)-glucans with a degree of polymerization from 2 to 7, act as primers and are elongated, a function analogous to that performed by glycogenin in glycogen synthesis.
MOS can originate from the activity of starch synthase III, starch phosphorylase, or starch debranching enzymes.
Being poorly water soluble, MOS seem able to evade the hydrolytic action of alpha-amylase (EC 3.2.1.1) and beta-amylase.[8]
Starch synthase and amylopectin synthesis
The synthesis of amylopectin requires the temporally coordinated action of at least four classes of enzymes: starch synthase isoenzymes, starch phosphorylase, starch branching enzymes (EC 2.4.1.18), and starch debranching enzymes.[2][4] It is also thought that, in many cases, these enzymes physically interact with each other to form multienzyme complexes, structures capable of increasing the efficiency of the metabolic pathway.[8]
It is generally accepted that starch granule growth occurs from a central core called the hilum, whose precise structure is unknown, although it appears to consist of a disordered arrangement of α-glucans.[9] The initiation of the hilum, the formation of normal starch granule morphology, and the degree of starch accumulation require the presence of SSIV, although SSIII and SSV are also thought to contribute, overlapping in function with SSIV.[10]
GBSS and amylose synthesis
Granule-bound starch synthase is involved in the synthesis of amylose.
This enzyme was first described by Luis Federico Leloir in the early 1960s, the same researcher who, in 1948, had discovered the main pathway for galactose metabolism, known as the Leloir pathway.[11]
Its catalytic action is not entirely simultaneous with that of the other starch synthases, as it requires the presence of an amylopectin matrix.[3]
In grasses, granule-bound starch synthase is present in two isoforms, encoded by distinct genes, referred to as GBSSI and GBSSII.[12]
GBSS requires, for its catalytic activity, the presence of a protein from the PTST family, PTST1, which enables its binding to the starch granule and whose action appears to be more important in chloroplasts than in amyloplasts.
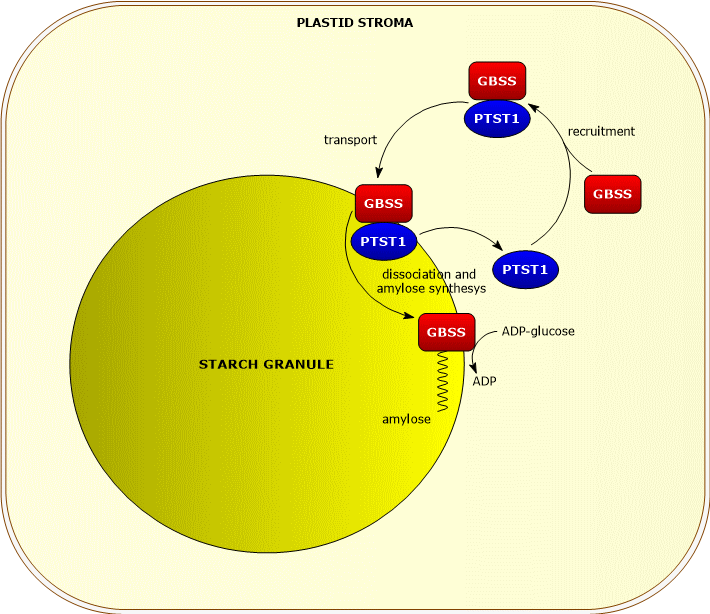
PTST1 appears to associate, in the plastid stroma, with GBSS. The complex binds to the nascent starch granule; the protein then dissociates from the enzyme, which begins catalyzing the elongation of malto-oligosaccharides, while the protein returns to the stroma to recruit another GBSS molecule.[13]
References
- ^ Zeeman S.C., Kossmann J., Smith A.M. Starch: its metabolism, evolution, and biotechnological modification in plants. Annu Rev Plant Biol 2010;61:209-34. doi:10.1146/annurev-arplant-042809-112301
- ^ a b Qu J., Xu S., Zhang Z., Chen G., Zhong Y., Liu L., Zhang R., Xue J., Guo D. Evolutionary, structural and expression analysis of core genes involved in starch synthesis. Sci Rep 2018;8(1):12736. doi:10.1038/s41598-018-30411-y
- ^ a b c Pfister B., Zeeman S.C. Formation of starch in plant cells. Cell Mol Life Sci 2016;73(14):2781-807. doi: 10.1007/s00018-016-2250-x
- ^ a b Crofts N., Abe N., Oitome N.F., Matsushima R., Hayashi M., Tetlow I.J., Emes M.J., Nakamura Y., Fujita N. Amylopectin biosynthetic enzymes from developing rice seed form enzymatically active protein complexes. J Exp Bot 2015;66(15):4469-82. doi:10.1093/jxb/erv212
- ^ Cuesta-Seijo J.A., Ruzanski C., Krucewicz K., Meier S., Hägglund P., Svensson B., Palcic M.M. Functional and structural characterization of plastidic starch phosphorylase during barley endosperm development. PLoS One 2017;12(4):e0175488. doi:10.1371/journal.pone
- ^ Abt M.R., Pfister B., Sharma M., Eicke S., Bürgy L., Neale I., Seung D., Zeeman S.C. STARCH SYNTHASE5, a noncanonical starch synthase-like protein, promotes starch granule initiation in Arabidopsis. Plant Cell 2020;32(8):2543-2565. doi:10.1105/tpc.19.00946
- ^ Seung D., Boudet J., Monroe J., Schreier T.B., David L.C., Abt M., Lu K.J., Zanella M., Zeeman S.C. Homologs of PROTEIN TARGETING TO STARCH control starch granule initiation in Arabidopsis leaves. Plant Cell 2017;29(7):1657-1677. doi:10.1105/tpc.17.00222
- ^ a b c Tetlow I.J., Bertoft E. A review of starch biosynthesis in relation to the building block-backbone model. Int J Mol Sci 2020;21(19):7011. doi:10.3390/ijms21197011
- ^ Ziegler G.R., Creek J.A., Runt J. Spherulitic crystallization in starch as a model for starch granule initiation. Biomacromolecules 2005;6(3):1547-54. doi:10.1021/bm049214p
- ^ Szydlowski N., Ragel P., Raynaud S., Lucas M.M., Roldán I., Montero M., Muñoz F.J., Ovecka M., Bahaji A., Planchot V., Pozueta-Romero J., D’Hulst C., Mérida A. Starch granule initiation in Arabidopsis requires the presence of either class IV or class III starch synthases. Plant Cell 2009;21(8):2443-57. doi:10.1105/tpc.109.066522
- ^ Leloir L.F., de Fekete M.A., Cardini C.E. Starch and oligosaccharide synthesis from uridine diphosphate glucose. J Biol Chem 1961;236:636-41. doi:10.1016/S0021-9258(18)64280-2
- ^ Vrinten P.L., Nakamura T. Wheat granule-bound starch synthase I and II are encoded by separate genes that are expressed in different tissues. Plant Physiol 2000;122(1):255-64. doi:10.1104/pp.122.1.255
- ^ Seung D., Soyk S., Coiro M., Maier B.A., Eicke S., and Zeeman S.C. PROTEIN TARGETING TO STARCH is required for localising GRANULE-BOUND STARCH SYNTHASE to starch granules and for normal amylose synthesis in Arabidopsis. PLoS Biol 2015;13:e1002080. doi:10.1371/journal.pbio.1002080