Starch phosphorylase, also known as alpha-glucan phosphorylase (EC 2.4.1.1), is a multimeric protein both with enzymatic and regulatory activity.
It plays an important role in carbohydrate metabolism in both prokaryotes and eukaryotes.[1]
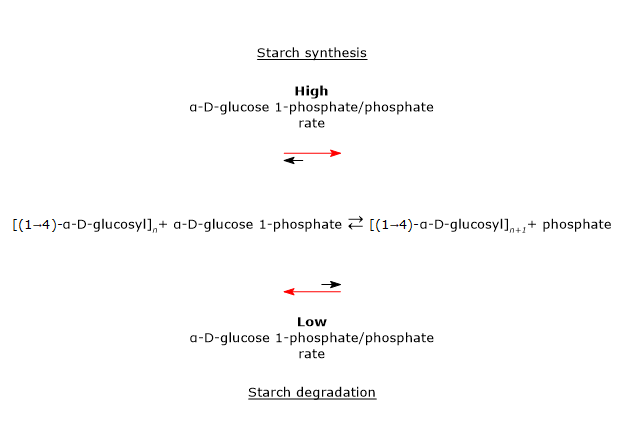
The enzyme catalyzes the transfer of a glucosyl unit from glucose 1-phosphate to the non-reducing end of a nascent α-(1→4)-glucan, forming an α-(1→4) glycosidic linkage.[2] This reaction is reversible, and its direction depends on the phosphate/glucose 1-phosphate ratio present in vivo.[3]
 The enzyme belongs to the family of glucosyltransferases (EC 2.4), along with other enzymes such as starch synthase (EC 2.4.1.21), involved in the synthesis of amylose and amylopectin, the polysaccharides that compose starch granules, glycogen phosphorylase (EC 2.4.1.1), involved in glycogenolysis, and glycogen synthase (EC 2.4.1.11), which participates in glycogen synthesis.[1]
The enzyme belongs to the family of glucosyltransferases (EC 2.4), along with other enzymes such as starch synthase (EC 2.4.1.21), involved in the synthesis of amylose and amylopectin, the polysaccharides that compose starch granules, glycogen phosphorylase (EC 2.4.1.1), involved in glycogenolysis, and glycogen synthase (EC 2.4.1.11), which participates in glycogen synthesis.[1]
It is important to note that while starch synthase uses ADP-glucose as the glucosyl donor and glycogen synthase uses UDP-glucose, starch phosphorylase utilizes glucose 1-phosphate.[4]
Additionally, starch phosphorylase appears to be involved in both the synthesis and degradation of amylose and amylopectin.[2]
In industrial applications, the phosphorolytic action of starch phosphorylase is employed in the production of glucose 1-phosphate and carbohydrates such as glucans and modified starches.[5]
Contents
Isoforms
At least two isoforms of starch phosphorylase are present in plants: one located in the stroma of plastids, named Pho1, and another with cytosolic localization, known as Pho2. Both isoforms play a critical role in the synthesis and degradation of starch.[6]
Starch degradation
Although alpha-amylase (EC 3.2.1.1) is the first enzyme to act in the polysaccharide degradation during the early stages of germination, and beta-amylase (EC 3.2.1.2) initiates transient starch degradation in chloroplasts, other enzymatic activities are also involved. These include alpha-glucan water dikinase (EC 2.7.9.4), phospho-glucan water dikinase (EC 2.7.9.5), and debranching enzymes.[7]
Of the two isoenzymes of starch phosphorylase, Pho1 appears to have an indirect or regulatory role, influencing the activity of the other enzymes involved in starch degradation.[6] In contrast, Pho2 is capable of directly degrading starch granules and other branched glucans.[8]
Starch synthesis
During starch biosynthesis, starch phosphorylase, particularly Pho1, appears to play both enzymatic and regulatory roles.
- Pho1 seems to be involved in the early stages of starch synthesis, contributing to the elongation of the nascent glucan chain.[4]
- Starch synthesis involves at least five classes of enzymes: ADP-glucose pyrophosphorylase (EC 2.7.7.27), starch synthases, starch branching enzymes (EC 2.4.1.18), starch debranching enzymes (EC 3.2.1.41), and starch phosphorylases.[9] In addition to these, there are also non-catalytic proteins essential for the correct assembly of the starch granule.[10]
In the endosperm of cereals, the formation of multienzyme complexes between starch synthases and branching enzymes depends not only on specific phosphorylation enents but also on the presence of Pho1.[11] - Starch phosphorylase is capable of forming a complex with disproportionating enzyme (EC 2.4.1.25).[11] This complex appears to synthesize short malto-oligosaccharides (MOS), which are α-(1→4)-glucans with a degree of polymerization between 2 and 7.[12][13]
MOS serve as primers for starch synthase IV and granule-bound starch synthase in the initial steps of amylopectin and amylose synthesis, respectively, role analogous to that of glycogenin in glycogen biosynthesis.[14]
MOS can also originate from the activity of starch debranching enzymes during the trimming of amylopectin molecules.[13]
References
- ^ a b Rathore R.S., Garg N., Garg S., Kumar A. Starch phosphorylase: role in starch metabolism and biotechnological applications. Crit Rev Biotechnol 2009;29(3):214-24. doi:10.1080/07388550902926063
- ^ a b Tickle P., Burrell M.M., Coates S.A., Emes M.J., Tetlow I.J., Bowsher C.G. Characterization of plastidial starch phosphorylase in Triticum aestivum L. endosperm. J Plant Physiol 2009;166(14):1465-78. doi:10.1016/j.jplph.2009.05.004
- ^ Newgard C.B., Hwang P.K., Fletterick R.J. The family of glycogen phosphorylases: structure and function. Crit Rev Biochem Mol Biol 1989;24(1):69-99. doi:10.3109/10409238909082552
- ^ a b Cuesta-Seijo J.A., Ruzanski C., Krucewicz K., Meier S., Hägglund P., Svensson B., Palcic M.M. Functional and structural characterization of plastidic starch phosphorylase during barley endosperm development. PLoS One 2017;12(4):e0175488. doi:10.1371/journal.pone
- ^ Kadokawa J.I. α-Glucan phosphorylase-catalyzed enzymatic reactions using analog substrates to synthesize non-natural oligo- and polysaccharides. Catalysts 2018;8(10):473. doi:10.3390/catal8100473
- ^ a b Yu G., Shoaib N., Xie Y., Liu L., Mughal N., Li Y., Huang H., Zhang N., Zhang J., Liu Y., Hu Y., Liu H., Huang Y. Comparative study of starch phosphorylase genes and encoded proteins in various Monocots and Dicots with emphasis on maize. Int J Mol Sci 2022;23:4518. doi:10.3390/ijms23094518
- ^ Fettke J., Eckermann N., Kötting O., Ritte G., Steup M. Novel starch-related enzymes and carbohydrates. Cell Mol Biol (Noisy-le-grand) 2007;52 Suppl:OL883-904.
- ^ Steup M., Robenek H., Melkonian M. In-vitro degradation of starch granules isolated from spinach chloroplasts. Planta 1983;158(5):428-36. doi:10.1007/BF00397736
- ^ Crofts N., Nakamura Y., Fujita N. Critical and speculative review of the roles of multi-protein complexes in starch biosynthesis in cereals. Plant Sci 2017;262:1-8. doi:10.1016/j.plantsci.2017.05.007
- ^ Seung D., Soyk S., Coiro M., Maier B.A., Eicke S., Zeeman S.C. PROTEIN TARGETING TO STARCH is required for localising GRANULE-BOUND STARCH SYNTHASE to starch granules and for normal amylose synthesis in Arabidopsis. PLOS Biol 2015;13(2):e1002080. doi:10.1371/journal.pbio.1002080
- ^ a b Crofts N., Abe N., Oitome N.F., Matsushima R., Hayashi M., Tetlow I.J., Emes M.J., Nakamura Y., Fujita N. Amylopectin biosynthetic enzymes from developing rice seed form enzymatically active protein complexes. J Exp Bot 2015;66(15):4469-82. doi:10.1093/jxb/erv212
- ^ Hwang S.K., Koper K., Satoh H., Okita T.W. Rice endosperm starch phosphorylase (Pho1) assembles with disproportionating enzyme (Dpe1) to form a protein complex that enhances synthesis of malto-oligosaccharides. J Biol Chem 2016;291(38):19994-20007. doi:10.1074/jbc.M116.735449
- ^ a b Tetlow I.J., Bertoft E. A review of starch biosynthesis in relation to the building block-backbone model. Int J Mol Sci 2020;21(19):7011. doi:10.3390/ijms21197011
- ^ Pfister B., Zeeman S.C., Rugen M.D., Field R.A., Ebenhöh O., Raguin A. Theoretical and experimental approaches to understand the biosynthesis of starch granules in a physiological context. Photosynth Res 2020;145:55-70. doi:10.1007/s11120-019-00704-y