Lipolysis is defined as the biochemical pathway responsible for the hydrolysis of triacylglycerols (TAGs) or triglycerides into non-esterified fatty acids (NEFAs) and glycerol.
The involved enzymes are called lipases.
This pathway is essential because triacylglycerols in their unhydrolyzed form cannot enter cells, as demonstrated first by Whitehead in 1909, nor exit. And in vertebrates there are three processes in which lipolysis is needed for the normal absorption or release of fatty acids and glycerol from the cells, and therefore, for lipid and energy homeostasis.
- Gastrointestinal lipolysis is responsible for the catabolism and the subsequent absorption of dietary triacylglycerols. The involved enzymes are lingual, gastric and pancreatic lipases, and pancreatic lipase related proteins 1, and 2 (PLRP1 and PLRP2).
- Vascular lipolysis mediates the hydrolysis of lipoprotein-associated triacylglycerols in the capillary bed. The involved enzymes are lipoprotein lipase (LPL), and hepatic lipase.
- Intracellular lipolysis is responsible for the hydrolysis of triacylglycerols stored in intracellular lipid droplets.
It involves neutral and acid lipases.
The acidic lipases have a pH optimum between 4 and 5.
They are the most important acid triacylglycerol hydrolase in lysosomes, and can hydrolyze cholesterol esters as well. These enzymes are thought to act primarily on lipoprotein-associated lipids, following their receptor-mediated endocytosis and sorting to lysosomes. However, their action is also associated with macroautophagy, a lysosomal pathway that catabolizes cytoplasmic inclusions such as aggregates of misfolded proteins, as well as damaged and superfluous organelles, releasing into the cytosol the hydrolysis products.
The cytosolic neutral lipases, with a pH optimum around 7.
They include:
adipose triglyceride lipase (ATGL);
hormone-sensitive lipase (HSL, EC 3.1.1.79);
monoacylglycerol lipase (MGL).
In the remainder of this article, we will analyze the intracellular lipolysis by the aforementioned neutral lipase and its hormonal and non-hormonal regulation, with particular attention to white adipose tissue.
Contents
- ATGL, HSL, MGL and lipolysis
- Hormonal regulation in white adipose tissue
- Non-hormonal regulation in white adipose tissue
- ATGL-mediated lipolysis in non-adipose tissues
- References
ATGL, HSL, MGL and lipolysis
Fatty acids deposited in the white adipose tissue as triacylglycerols represent the largest energy store in higher eukaryotes.
When energy demand increases, such as during intense and prolonged physical activity, triacylglycerol hydrolysis occurs and fatty acids are released into the blood.
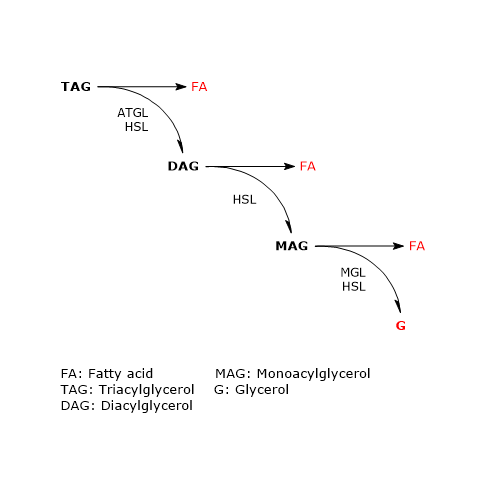
In adipose tissue the sequential action of these three enzymes leads to complete hydrolysis of triacylglycerols. In this process, both in vivo and in cultured adipocytes, ATGL and HSL account for more than 90 percent of the lipolytic activity.
Adipose triglyceride lipase
Of the three aforementioned neutral lipases, ATGL is the more recently discovered.
Its role in triacylglycerol catabolism became clear after studies conducted on mutant mice lacking the enzyme, and on humans with mutations in the gene encoding it. In humans, for example, a systemic triacylglycerol accumulation, and therefore of body fat, and cardiomyopathy has been observed. This clearly indicates that ATGL activity is required for efficient mobilization of triacylglycerols in adipose and non-adipose tissues.
The human enzyme, encoded by a gene on chromosome 11p15.5, belongs to the family of proteins containing a patatin domain, which includes 8 murine and 9 human members. The patatin domain is present in the N-terminal half of the protein and also contains the active site of the enzyme. Conversely, the C-terminal half has primarily a regulatory function, and also includes the hydrophobic region responsible for the binding of lipid droplets, essential for in vivo enzyme activity.
Orthologous enzymes are present in essentially all eukaryotes, including invertebrates, vertebrates, as well as fungi and plants.
The enzyme catalyzes the first step in triacylglycerol hydrolysis, leading to the formation of diacylglycerols (DAG) or diglycerides, and fatty acids:
Triacylglycerol + H20 → Diacylglycerols + Fatty acids
ATGL preferentially hydrolyzes sn-2 ester bonds, but as a consequence of the interaction with CGI-58, its selectivity broadens to the sn-1 bond.
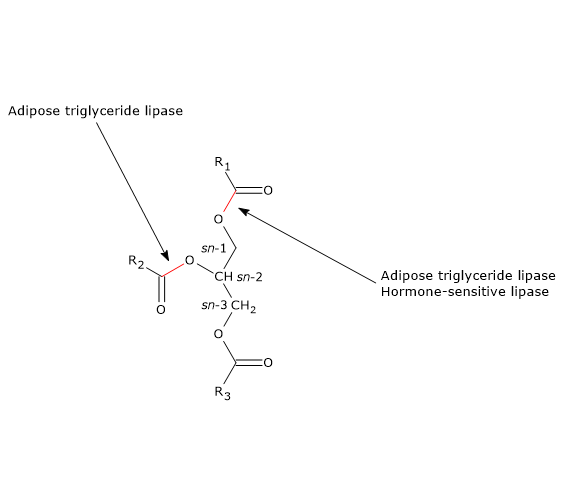
Therefore, ATGL has a central role in the metabolism, as also suggested by studies in fasted mutant mice lacking the enzyme, in which the lack of non-esterified fatty acids causes a high glucose consumption for energy purposes; and hypoglycaemia, hypometabolism and hypothermia occur with fasting for more than 6 hours.
Compared to the activity towards triacylglycerols, the enzyme exhibits minor or no catalytic activity towards monoacylglycerols (MAGs) or monoglycerides, diacylglycerols, cholesterol esters, and retinol esters.
Enzyme activity is subject to regulation by interacting with activating or inhibitory proteins, some of which are localized on lipid droplets, and described below.
CGI-58
Like pancreatic lipase and LPL, which are much more active in the presence of protein coactivators, ATGL catalytic activity is increased by the activator protein comparative gene identification-58 (CGI-58), which therefore stimulates the first step of intracellular lipolysis.
It is a highly conserved protein among species, encoded in humans by a gene on chromosome 3p21. It interacts with the patatin domain of ATGL. The maximal stimulation occurs at approximately equimolar concentrations of the two proteins.
The importance of its stimulatory action is underlined by the fact that its deficiency or malfunctioning results in a severe systemic accumulation of triacylglycerols both in men and mice.
CGI-58 is regulated primarily by its interaction with perilipin-1, a protein that covers lipid droplets. CGI-58, at least in vitro, also acts as an acyl-CoA-dependent acylglycerol-3-phosphate acyltransferase.
G0S2
It is an inhibitor of ATGL, originally identified in mononuclear blood cells, where it acts at the G0 to G1 transition of the cell cycle, and therefore called G0G1 switch protein 2 (G0S2).
In humans, it is encoded by a gene on chromosome 1q32.2.
It is present in many tissues, with the highest levels in adipose tissue and liver, followed by ovary, muscle, and kidney. It is found in different cellular compartments, such as cytoplasm, mitochondria, endoplasmic reticulum, and lipid droplets. These different cellular locations may reflect the different functions that the protein performs, such as the regulation of:
- lipolysis;
- cell cycle;
- possibly apoptosis, through its ability to interact with Bcl2, a mitochondrial antiapoptotic factor.
G0S2, like CGI-58, interacts with the patatin domain of ATGL, and, at least in vitro, lipid droplet binding and enzyme inhibition depend on physical interaction between the N-terminal region of G0S2 and the patatin domain of the enzyme. However, it does not seem that this interaction directly competes with the binding of the activator CGI-58.
PEDF
Another protein involved in the regulation of ATGL activity seems to be pigment epithelium derived factor (PEDF), which induces the hydrolysis of triacylglycerols in adipose tissue, liver, and muscle via lipase activity.
It is a widely expressed protein, belonging to the noninhibitory Serpin family, and with a large spectrum of activities, such as anti-inflammatory, antioxidative, neuroprotective, antitumorigenic , and antiangiogenic effects. The protein binds to the enzyme and activates it. Its activity may be involved in the development of hepatic steatosis and the pathogenesis of insulin resistance.
Finally, it seems that ATGL delivery to lipid droplets requires vesicular transport. In fact, its translocation to the lipid droplets is blocked in the absence of Sar1, ARF1 or GBF1, which are protein components of the transport machinery; and the enzyme remains associated with the endoplasmic reticulum, from which lipid droplets are believed to bud off.
Hormone-sensitive lipase
In the early sixties of last century it was noted that lipolytic activity in adipose tissue was induced by hormones. In 1964, both hormone-sensitive lipase and monoacylglycerol lipase of adipose tissue were isolated and characterized.
It was immediately clear that HSL was more efficient as a diacylglycerol hydrolase than triacylglycerol hydrolase, in vitro by a factor of 10-fold. Despite this, it was assumed that HLS was the rate-limiting enzyme in the catabolism of triacylglycerols in adipose and many other tissues. Many years later, in 2000, it was however observed that enzyme-deficient mice showed no signs of triacylglycerol accumulation in adipose and other tissues, whereas they accumulated large quantities of diacylglycerols in many tissues. This suggested that HLS was more important as diacylglycerol hydrolase than as triacylglycerol hydrolases, which is now commonly accepted.
Diacylglycerols + H20 → Monoacylglycerols + Fatty acids
However, it is a multifunctional enzyme able to hydrolyze, in addition to diacylglycerols in which it has a stereo-preference for sn-3 ester bonds, and triacylglycerols, in which it preferentially hydrolyses sn-1 ester bond, also the ester bonds of other lipids, such as monoacylglycerols, retinyl esters, and cholesteryl esters.
HSL is encoded by a gene on chromosome 19q13.2. Alternative splicing leads to significant variations in the 5′ region of the transcripts, and therefore tissue specific mRNAs and proteins of different sizes.
The enzyme expression profile is essentially similar to that of ATGL. The highest abundance of mRNA and protein is found in white and brown adipose tissue; in many other tissues and cells, including muscle, pancreatic beta-cells, steroidogenic cells, and macrophages, HSL gene expression is low.
Unlike adipose triglyceride lipase, which has orthologous enzymes in all eukaryotes, hormone-sensitive lipase is less ubiquitous; for example, no orthologous proteins are known in birds, in D. melanogaster, C. elegans, and S. cerevisiae.
Finally, unlike ATGL, no mutations in HSL gene have been observed in humans.
Three functional regions been identified in the structure of the protein:
- an N-terminal domain, which is believed to mediate enzyme dimerization, lipid binding, and interaction with FABP4, a protein which increases HSL catalytic activity.
- A C-terminal domain, which harbors a structural fold common to many esterases and lipases, called the alpha/beta hydrolase fold; it contains the classical catalytic triad of human hydrolase, Ser424, Asp693, and His72, that is, the active site.
- The third region is the regulatory module of HSL. It is located within the catalytic domain and contains at least five phosphorylation sites, on as many serine residues, two of which, Ser650 and Ser663, seem to be particularly important for its activity.
Monoacylglycerol lipase
The protein is considered to be the rate-limiting enzyme for the catabolism of monoacylglycerols derived from hydrolysis of:
- plasma lipoprotein triacylglycerols by lipoprotein lipase;
- intracellular triacylglycerol by ATGL and HSL;
- intracellular phospholipids by phospholipase C and diglyceride lipase α and β.
Monoacylglycerols + H20 → Fatty acids + Glycerols
In humans, the enzyme is encoded by a gene on chromosome 3q21.3, and is ubiquitously expressed, with highest expression in adipose tissue; however, high expression levels are also found in hepatocytes and muscle cells.
The enzyme is localized on lipid droplets, cell membranes, and in cytoplasm.
The importance of MGL in the breakdown of monoacylglycerols has been confirmed by studies conducted on mutant mice: its lack impairs lipolysis and is associated with similar increases in the level of monoacylglycerols in adipose and non-adipose tissues.
Other enzymes with monoacylglycerol hydrolase activities are HSL and ABHD6.
The protein shares homology to lysophospholipases, esterases, and haloperoxidases.
It seems that neither mRNA concentration nor catalytic activity is regulated by hormones or cell energy charge.
Hormonal regulation in white adipose tissue
In white adipose tissue, triacylglycerol hydrolysis on the surface of lipid droplets is mainly regulated post-translationally by phosphorylations, protein-protein interactions, and translocations of the participating proteins.
Beta-adrenergic stimulation
Adrenaline and glucagon, as a result of binding to specific β-adrenergic receptors on adipocytes, activate ATGL and HSL, which hydrolyze triacylglycerols in a coordinated manner.
A central role in the process leading to enzyme activation is performed by perilipin-1, a lipid-droplet associated protein found only in cells which can be beta-adrenergically stimulated. As a result of such stimulation, perilipin-1 is phosphorylated on six serine residues by protein kinase A (PKA).
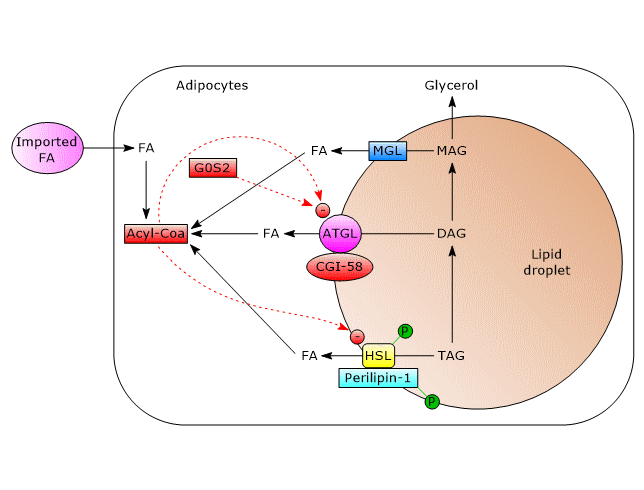
- ATGL
In unstimulated adipocytes, CGI-58 is localized on the surface of lipid droplets, mostly bound to perilipin-1, and does not interact with ATGL. In this state, ATGL activity, and therefore lipolysis, are low.
Phosphorylation of CGI-58 and perilipin-1 by PKA leads to the release of CGI-58 which in turn, in a not well understood way, actives the enzyme.
Even ATGL can be phosphorylated on at least two serine residues, Ser404 and Ser428, but not by PKA. However, the importance of these phosphorylations in the regulation of the enzyme activity is not yet clear. - HSL
In adipose tissue, its activity is strongly induced by the stimulation of beta-adrenergic receptors, whereas it is strongly inhibited by insulin.
Phosphorylation increases moderately, by about two-fold, enzyme activity. For complete activation, the enzyme must gain access to lipid droplets, which is mediated by phosphorylated perilipin-1. How?
In unstimulated cells, perilipin-1 is not phosphorylated and prevents HSL binding to lipid droplets. The phosphorylations allow the binding of HSL to perilipin-1, in particular to the N-terminal region of the protein, and therefore the access to lipid droplets.
Therefore, beta-adrenergic stimulation, via HSL phosphorylation and translocation to lipid droplet surface, leads to an increased activity of the enzyme in white adipose tissue of about 100-fold.
Other kinases, such as AMPK or Ca2+/calmodulin-dependent kinase also phosphorylate the enzyme to modulate its activity. - CGI-58
beta-adrenergic stimulation, like fasting, has a little effect on the expression of the protein in the adipose tissue.
Inhibition by insulin
Insulin mediates the deactivation of lipolysis by acting at transcriptional level, on the activity of ATGL and HSL, and via sympathetic nervous system.
- The hormone causes the transcriptional downregulation of ATGL and HSL gene expression.
- It induces the phosphorylation and activation of phosphodiesterase isoforms by PKB/AKT, cAMP hydrolysis by these phosphodiesterase, the consequent inactivation of PKA, and then the prevention of the phosphorylation of perilipin-1 and HSL.
- Insulin also inhibits lipolysis via a central mechanism that involves the sympathetic nervous system. In fact, increased hormone levels in the brain inhibit phosphorylation of perilipin-1 and HSL.
Furthermore, insulin also stimulates the expression of G0G2.
Non-hormonal regulation in white adipose tissue
Non-hormonal factors can also play a role in the regulation of triacylglycerol hydrolysis.
- Recent works have shown an inhibitory effect of long chain acyl-CoAs, such as palmitoyl-CoA and oleoyl-CoA, on ATGL activity.
Note: palmitic acid and oleic acid are two highly abundant fatty acids in white adipose tissue.
Similarly to what described for HSL, long chain acyl-CoAs act as non-competitive inhibitors. This inhibition may be an effective feedback mechanism for the control of triacylglycerol hydrolysis and the protection of the cells from lipotoxic concentrations of molecules such as diacylglycerol, acyl-CoAs, and ceramides. - Receptor-interacting protein 140 (RIP-140), as a result of an increase in cellular lipid content, induces lipolysis by binding to perilipin-1 on lipid droplets. Thanks to this interaction, perilipin-1 more efficiently recruits HSL to lipid droplets and enhances complex formation between ATGL and its activator CGI-58.
ATGL-mediated lipolysis in non-adipose tissues
ATGL and HSL are also expressed in most non-adipose tissues, although in low levels. This has raised the question whether the intervention of other lipases occur to ensure efficient lipolysis. For example, in the liver of fasted mice, ATGL accounts for less than 50 percent of neutral hydrolysis of triacylglycerols, an activity that is, however, important because its absence leads to the development of hepatosteatosis. But even in its absence, a good activity on triacylglycerols is retained in the hepatocytes. This ensures no apparent defect in VLDL production, whose assembly and secretion requires a considerable mobilization of hepatic triacylglycerol stores.
Anyway, in non-adipose tissues, such as skeletal muscle, heart, and liver, ATGL-mediated lipolysis follows a different mechanism.
Perilipin-5 replaces perilipin-1. During fasting, it recruits both CGI-58 and ATGL to lipid droplets by direct binding of the two proteins. The role of perilipin-5 in this complex is not yet clear, but it seems to be involved in the interaction of lipid droplets with mitochondria and the inhibition of ATGL-mediated triacylglycerol hydrolysis.
In skeletal muscle, hormone-sensitive lipase is activated by phosphorylation in response to muscle contraction and adrenaline.
In non-adipose tissues lacking perilipin-1 the role of HSL is less well characterized and the enzyme expression is low. The interaction between HSL and lipid droplets may involve perilipin-2 and perilipin-5.
References
- Boeszoermenyi A., Nagy H.M., Arthanari A., Pillip C.J., Lindermuth H., Eulogio Luna R., Wagner G., Zechner R., Zangger K., and Oberer M. Structure of a CGI-58 motif provides the molecular basis of lipid droplet anchoring. J Biol Chem 2015;290(44):26361-26372. doi:10.1074/jbc.M115.682203
- Ho P.C., Chuang Y.S., Hung C.H., and Wei L.N. Cytoplasmic receptor-interacting protein 140 (RIP140) interacts with perilipin to regulate lipolysis. Cell Signal 2011;23:1396-1403. doi:10.1016/j.cellsig.2011.03.023
- Lass A., Zimmermann R., Oberer M., Zechner R. Lipolysis – A highly regulated multi-enzyme complex mediates the catabolism of cellular fat stores. Prog Lipid Res 2011;50;14-27. doi:10.1016/j.plipres.2010.10.004
- Nagy H.M., Paar M., Heier C., Moustafa T., Hofer P., Haemmerle G., Lass A., Zechner R., Oberer M., Zimmermann R. Adipose triglyceride lipase activity is inhibited by long-chain acyl-coenzyme A. Biochim Biophys Acta 2014;1841:588-594. doi:10.1016/j.bbalip.2014.01.005
- Zechner R., Zimmermann R., Eichmann T.O., Kohlwein S.D., Haemmerle G., Lass A., and Madeo F. FAT SIGNALS – Lipases and lipolysis in lipid metabolism and signaling. Cell Metab 2012;15:279-291. doi:10.1016/j.cmet.2011.12.018